flim-imager-releases
FLIM Imager Data Export
Table of Contents
Introduction
FLIM Imager software allows seamless export of imaging analyzed data to JSON files, with convenient plotting and visualization capabilities. This guide provides an in-depth exploration of the JSON files structure, offering a comprehensive understanding of how exported data is formatted and can be leveraged.
Single-Frame Imaging File Format
Here a detailed explanation of the exported single-frame JSON data file structure:
Header (Metadata):
The file structure starts with the header field, which contains the following metadata info:
- type: Information about the file type. In this case, “Frame”. (string);
- file_id: ASCII values which uniquely identify the file type. In this case the matching string value is “IMF1”. (int[]);
- setup: The microscope setup used (Default or Abberior configuration);
- abberior_multichannel_assignment_mode: Information about the Abberior multi-channel imaging reconstruction assigment (Frame, Line or Pixel). This field is only present when in Abberior setup mode;
- channels: Information about enabled channels (boolean[]);
- laser_period_ns: Laser period in nanoseconds (float);
- step: Imaging type of experiment (Scouting/Imaging) (string);
- reconstruction: Imaging type of reconstruction (At the moment, only PLF/pixel-line-frame is supported) (string);
- image_width: Width of the acquired image in pixels (int);
- image_height: Height of the acquired image in pixels (int);
Data Records:
Following the header data, data field contains the decay values for each pixels in the frame, structured as a three-level array:
- 1st Level: An array containing one subarray related to the acquired data acquisition channel ;
- 2nd Level: The subarray corresponding to the data acquisition channel contains as many subarrays as there are total pixels in the acquired frame;
- 3rd Level: Each subarray corresponding to a pixel contains a variable number of subarrays, each with a length of 2. In these subarrays, the first index represents the bin index (pixels are binned into 256 bins, with photon counts calculated for each bin), and the second index represents the decay value for that bin. These subarrays have a variable length because bins with null photon counts are omitted for optimization purposes.
Cumulative Imaging File Format
Here a detailed explanation of the exported global image JSON data file structure:
Header (Metadata):
The file structure starts with the header field, which contains the following metadata info:
- type: Information about the file type. In this case, “Global”. (string);
- file_id: ASCII values which uniquely identify the file type. In this case the matching string value is “IMG1”. (int[]);
- setup: The microscope setup used (Default or Abberior configuration);
- abberior_multichannel_assignment_mode: Information about the Abberior multi-channel imaging reconstruction assigment (Frame, Line or Pixel). This field is only present when in Abberior setup mode;
- channels: Information about enabled channels (boolean[]);
- laser_period_ns: Laser period in nanoseconds (float);
- step: Imaging type of experiment (Scouting/Imaging) (string);
- reconstruction: Imaging type of reconstruction (At the moment, only PLF/pixel-line-frame is supported) (string);
- image_width: Width of the acquired image in pixels (int);
- image_height: Height of the acquired image in pixels (int);
- frames: Total number of acquired frames (int);
Data Records:
Following the header data, data field contains the decay values for each pixels in the frame, structured as a three-level array:
- 1st Level: An array containing as many subarrays as there are active acquisition channels;
- 2nd Level: Each subarray corresponding to an acquisition channel contains as many subarrays as there are total pixels in the acquired frame;
- 3rd Level: Each subarray corresponding to a pixel contains a variable number of subarrays, each with a length of 2. In these subarrays, the first index represents the bin index (pixels are binned into 256 bins, with photon counts calculated for each bin), and the second index represents the decay value for that bin. These subarrays have a variable length because bins with null photon counts are omitted for optimization purposes. In this case (Cumulative Imaging), photon counts are cumulative sum of each frames counts for each bin.
Single-Frame Phasors File Format
Here a detailed explanation of the exported single-frame phasors JSON data file structure:
Header (Metadata):
The file structure starts with the header field, which contains the following metadata info:
- type: Information about the file type (Frame). (string);
- file_id: ASCII values which uniquely identify the file type (“IPF1” for Single frame export). (int[]);
- setup: The microscope setup used (Default or Abberior configuration);
- abberior_multichannel_assignment_mode: Information about the Abberior multi-channel imaging reconstruction assigment (Frame, Line or Pixel). This field is only present when in Abberior setup mode;
- channels: Information about enabled channels (boolean[]);
- laser_period_ns: Laser period in nanoseconds (float);
- step: Imaging type of experiment (Scouting/Imaging) (string);
- reconstruction: Imaging type of reconstruction (At the moment, only PLF/pixel-line-frame is supported) (string);
- image_width: Width of the acquired image in pixels (int);
- image_height: Height of the acquired image in pixels (int);
- frames: Total number of acquired frames (int);
- tau_ns: tau (ns) value (float);
- harmonics: Number of phasor harmonics (int);
Data Records:
Following the header field, data field contains the following information:
- frame: the number of the last frame acquired (int);
- channel: the acquisition channel for which the phasor data was calculated (int);
- harmonic: the harmonic for which the phasor data was calculated (int);
- g_data: an array which contains as many subarray as there are pixels in the acquired frame/image. Each pixel sub-array contains the g coordinate for the phasor representation;
- s_data: an array which contains as many subarray as there are pixels in the acquired frame/image. Each pixel sub-array contains the s coordinate for the phasor representation;
Cumulative Phasors File Format
Here a detailed explanation of the exported global phasors JSON data file structure:
Header (Metadata):
The file structure starts with the header field, which contains the following metadata info:
- type: Information about the file type (Global). (string);
- file_id: ASCII values which uniquely identify the file type (“IPG1” for Global export). (int[]);
- setup: The microscope setup used (Default or Abberior configuration);
- abberior_multichannel_assignment_mode: Information about the Abberior multi-channel imaging reconstruction assigment (Frame, Line or Pixel). This field is only present when in Abberior setup mode;
- channels: Information about enabled channels (boolean[]);
- laser_period_ns: Laser period in nanoseconds (float);
- step: Imaging type of experiment (Scouting/Imaging) (string);
- reconstruction: Imaging type of reconstruction (At the moment, only PLF/pixel-line-frame is supported) (string);
- image_width: Width of the acquired image in pixels (int);
- image_height: Height of the acquired image in pixels (int);
- frames: Total number of acquired frames (int);
- tau_ns: tau (ns) value (float);
- harmonics: Number of phasor harmonics (int);
Data Records:
Following the header field, phasors_data array contains dictionaries with the following information:
- frame: the number of the last frame acquired (int);
- channel: the acquisition channel for which the phasor data was calculated (int);
- harmonic: the harmonic for which the phasor data was calculated (int);
- g_data: an array which contains as many subarray as there are pixels in the acquired image. Each pixel sub-array contains the g coordinate for the phasor representation;
- s_data: an array which contains as many subarray as there are pixels in the acquired image. Each pixel sub-array contains the s coordinate for the phasor representation;
Following phasors_data , intensities_data field contains the decay values for each pixels in the acquired image, structured as a three-level array:
- 1st Level: An array containing one subarray corresponding to the current active acquisition channel data ;
- 2nd Level: The active channel subarray contains as many subarrays as there are total pixels in the acquired image;
- 3rd Level: Each subarray corresponding to a pixel contains a variable number of subarrays, each with a length of 2. In these subarrays, the first index represents the bin index (pixels are binned into 256 bins, with photon counts calculated for each bin), and the second index represents the decay value for that bin. These subarrays have a variable length because bins with null photon counts are omitted for optimization purposes. In this case (Cumulative analysis), photon counts are cumulative sum of each frames counts for each bin.
Data Visualization
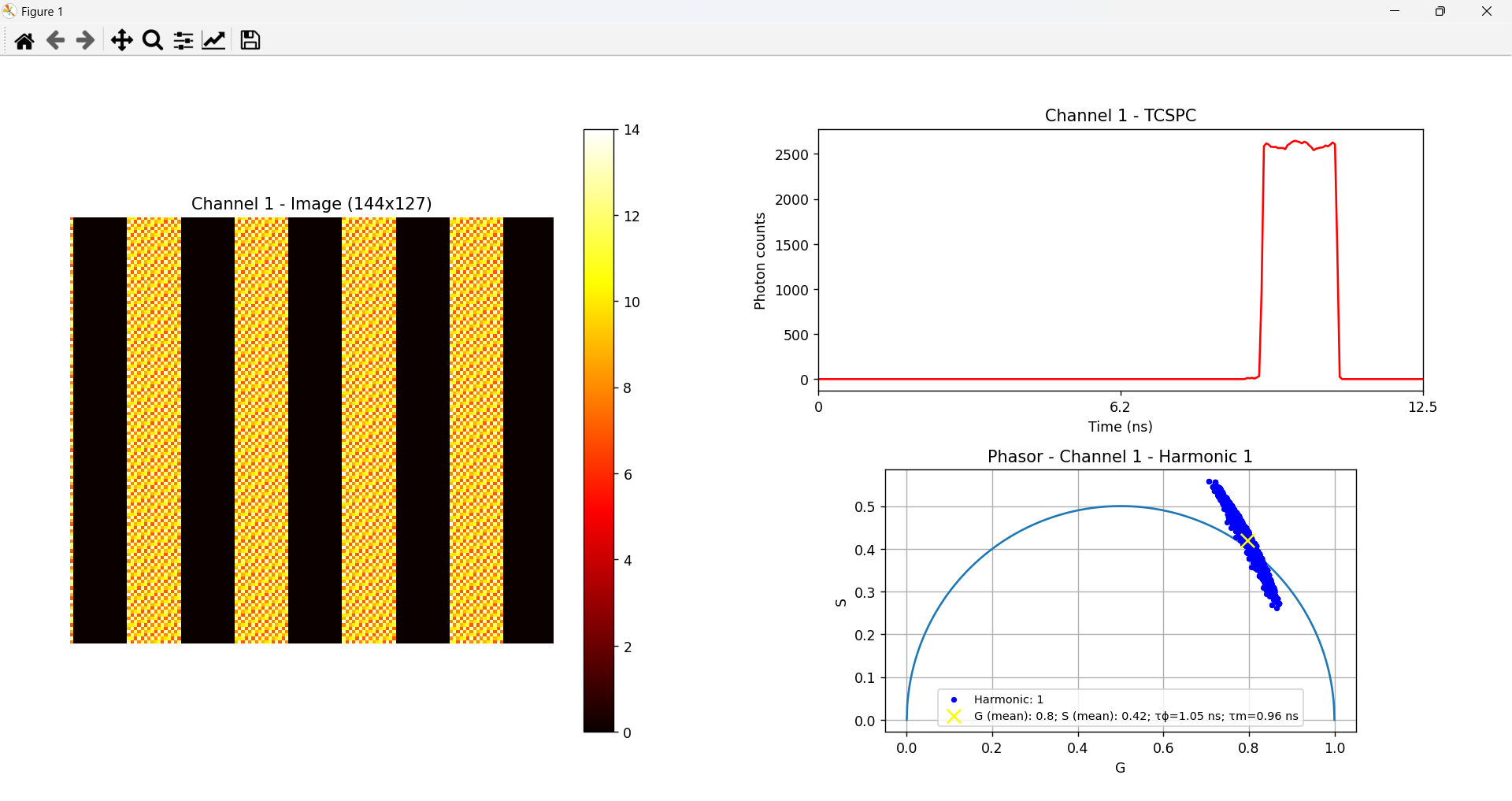
The script files are automatically downloaded along with the acquisition .bin file once the acquisition is complete and a file name and file path has been chosen. When running the provided Python or Matlab scripts, the image reconstruction is performed. This process generates a visual representation of the acquired data based on the specified colormap. Along with the image, the script also produces the corresponding global TCSPC plot, which visualizes the time-correlated single-photon counting data across the entire sample. If the data is exported in FLIM Phasors Mode, the output also includes a phasor plot representing the frame or the globally acquired image, providing an additional graphical analysis of the lifetime distribution.
Follow the guide below if you wish to use the Python script:
-
Python script:
-
Open the terminal and navigate to the directory where the saved files are located (it is advisable to save and group them in a folder):
cd YOUR_DOWNLOADED_DATA_ROOT_FOLDER - Create a virtual environment using the command:
python -m venv venv - Activate the virtual environment with the command:
venv\Scripts\activate - Install the necessary dependencies listed in the automatically downloaded requirements.txt with:
pip install -r requirements.txt - Run your script with:
python YOUR_SCRIPT_NAME.py
-
-
Matlab script:
Simply open your MATLAB command window prompt and, after navigating to the folder containing the script, type the name of the script to launch it.
Script Configuration
In the provided Python and Matlab scripts, users will find an initial section of the code containing configurable variables along with commented instructions explaining their function. This section allows users to adjust key parameters for analysis. Depending on the script type users can find:
- file_path: This variable lets users specify the imaging .JSON file to analyze. If the user wishes to change the file being processed, they can modify this variable to point to a different file.
- imaging_file_path (only in the single-frame phasors script): This variable lets users specify the imaging .JSON file to analyze. If the user wishes to change the file being processed, they can modify this variable to point to a different file.
- phasors_file_path (only in the phasors script): This variable lets users specify the phasors .JSON file to analyze. If the user wishes to change the file being processed, they can modify this variable to point to a different file.
- channel: Users can select which of the active channels to visualize during analysis. By default, this is set to the first active channel, but it can be changed to any other active channel.
- colormap: This variable allows users to set the colormap used in the image reconstruction. By default, the colormap is set to hot.
- frame (only in the single-frame reconstruction script): For frame-specific analysis, users can select which frame to analyze by specifying the frame number.
- harmonic (only in the phasors script): users can select which phasor harmonic to analyze by specifying the harmonic number.
Useful Links
To view the detailed JSON data schemas for each exported file type, download this ZIP.
For more details about the project follow these links:
License
Distributed under the MIT License.
Contact
FLIM LABS: info@flimlabs.com
Project Link: FLIM Imager
